Topological data analysis
Topological data analysis is a new area of study aimed at having applications in areas such as data mining and computer vision. The main problems are:
- how one infers high-dimensional structure from low-dimensional representations; and
- how one assembles discrete points into global structure.
The human brain can easily extract global structure from representations in a strictly lower dimension, i.e. we infer a 3D environment from a 2D image from each eye. The inference of global structure also occurs when converting discrete data into continuous images, e.g. dot-matrix printers and televisions communicate images via arrays of discrete points.
The main method used by topological data analysis is:
- Replace a set of data points with a family of simplicial complexes, indexed by a proximity parameter.
- Analyse these topological complexes via algebraic topology — specifically, via the new theory of persistent homology.
- Encode the persistent homology of a data set in the form of a parameterized version of a Betti number which will be called a barcode.
Contents |
Point cloud data
Data is often represented as points in a Euclidean n-dimensional space En. The global shape of the data may provide information about the phenomena that the data represent.
One type of data set for which global features are certainly present is the so-called point cloud data coming from physical objects in 3D. E.g. a laser can scan an object at a set of discrete points and the cloud of such points can be used in a computer representation of the object. Point cloud data refers to any collection of points in En or a (perhaps noisy) sample of points on a lower-dimensional subset.
For point clouds in low-dimensional spaces there are numerous approaches for inferring features based on planar projections in the fields of computer graphics and statistics. Topological data analysis is needed when the spaces are high-dimensional or too twisted to allow planar projections.
To convert a point cloud in a metric space into a global object, use the point cloud as the vertices of a graph whose edges are determined by proximity, then turn the graph into a simplicial complex and use algebraic topology to study it. An alternative approach is the minimum spanning tree-based method in the geometric data clustering.[1] If a group of data points forms a cluster, then the geometry of this point cloud can be determined.
Persistent homology
See homology for an introduction to the notation.
Persistent homology essentially calculates homology groups at different resolutions to see which features persist for long periods of time. It is assumed that important features and structures are the ones that persist. We define persistent homology as follows:
Let 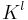 be a filtration. The p-persistent kth homology group of
be a filtration. The p-persistent kth homology group of 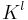 is
is 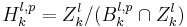 .
.
If we let 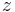 be a nonbounding
be a nonbounding 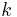 -cycle created at time
-cycle created at time 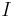 by simplex
by simplex 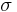 and let
and let 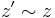 be a homologous
be a homologous 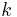 -cycle that becomes a boundary cycle at time
-cycle that becomes a boundary cycle at time 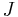 by simplex
by simplex 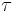 , then we can define the persistence interval associated to
, then we can define the persistence interval associated to 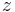 as
as 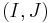 . We call
. We call 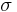 the creator of
the creator of 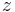 and
and 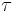 the destroyer of
the destroyer of 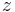 . If
. If 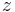 does not have a destroyer, its persistence is
does not have a destroyer, its persistence is 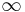 .
.
Instead of using an index-based filtration, we can use a time-based filtration. Let 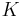 be a simplicial complex and
be a simplicial complex and 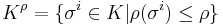 be a filtration defined for an associated map
be a filtration defined for an associated map 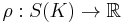 that maps simplices in the final complex to real numbers. Then for all real numbers
that maps simplices in the final complex to real numbers. Then for all real numbers 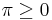 , the
, the 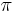 -persistent kth homology group of
-persistent kth homology group of 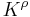 is
is 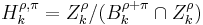 . The persistence of a
. The persistence of a 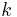 -cycle created at time
-cycle created at time 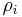 and destroyed at
and destroyed at 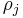 is
is 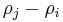 . [2]
. [2]
There are various software packages for computing persistence intervals of a finite filtration, such as jPlex, Dionysus and the Perseus software projects.
See also
- Dimensionality reduction
- Data mining
- Computer vision
- Computational topology
- Digital topology
- Digital Morse theory
- Shape analysis
- Size theory
- Structured data analysis (statistics)
References
- ^ C. T. Zahn (1971): "Graph-theoretical methods for detecting and describing gestalt clusters", IEEE Transactions on Computers, pp. 68-86, Vol. 20, No. 1
- ^ Afra J. Zomorodian (2005): Topology for Computing. Cambridge Monographs on Applied and Computational Mathematics.
Further reading
- Topological Methods in Scientific Computing, Statistics and Computer Science Stanford group
- BARCODES: THE PERSISTENT TOPOLOGY OF DATA
- Topological Data Analysis: the algebraic topology of point data clouds?
- Sanjay Rana (2004). Topological Data Structures for Surfaces. John Wiley and Sons. ISBN 978-0470851517. http://books.google.com/books?id=bfPWmkohAQoC.
- TOPOLOGY AND DATA, GUNNAR CARLSSON, BULLETIN (New Series) OF THE AMERICAN MATHEMATICAL SOCIETY, Volume 46, Number 2, April 2009, Pages 255–308, Article electronically published on January 29, 2009
- Computational Topology: An Introduction, Herbert Edelsbrunner, John L. Harer, AMS Bookstore, 2010, ISBN 9780821849255
- Topological Methods in Data Analysis and Visualization: Theory, Algorithms, and Applications, Editors Valerio Pascucci, Hans Hagen, Xavier Tricoche, Julien Tierny, Springer, 2010, ISBN 9783642150135
- Weinberger, Shmuel (2011), "What Is . . . Persistent Homology?", AMS Notices 58 (01): 36–39, http://www.ams.org/notices/201101/rtx110100036p.pdf.